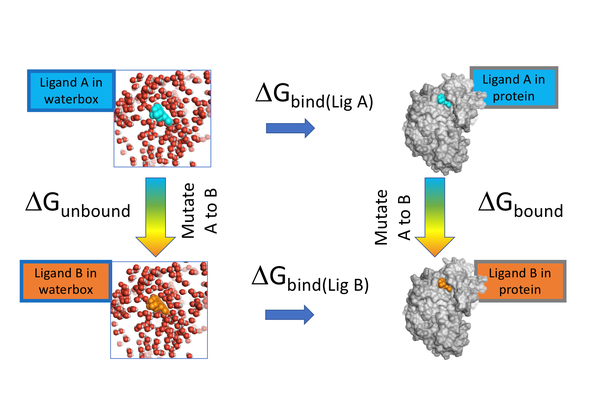
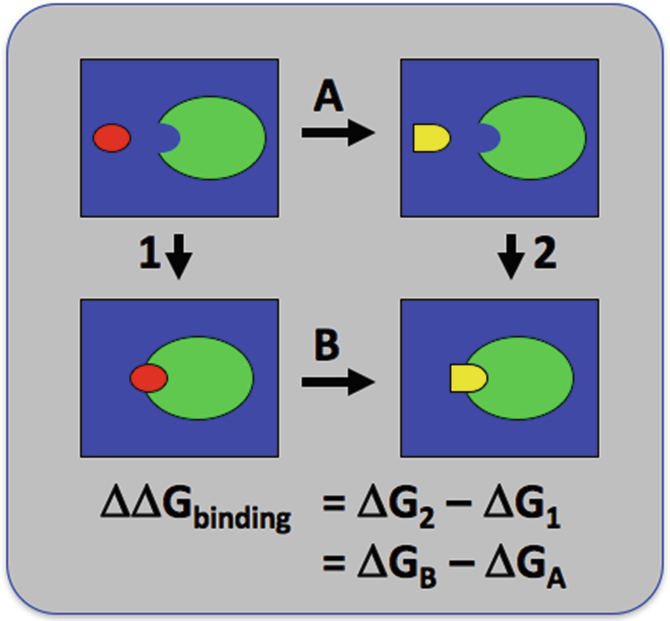
FEP calculation of the relative binding free energy due to a mutation.... | Download Scientific Diagram
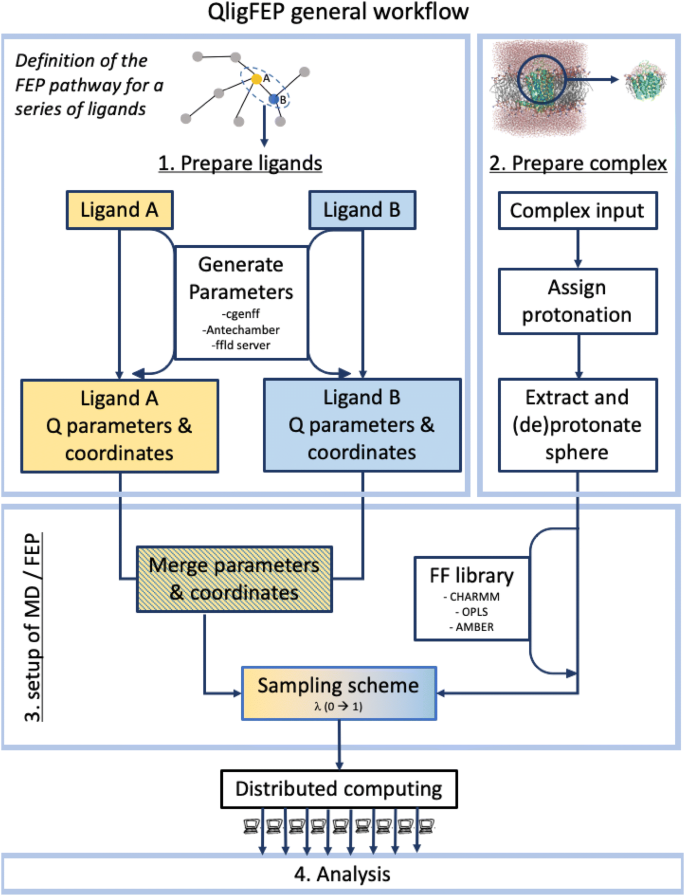
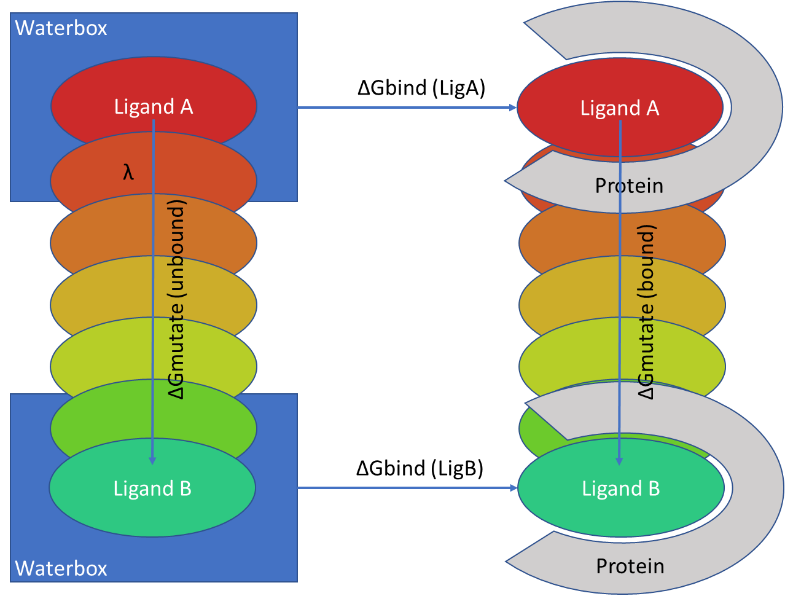
QligFEP: an automated workflow for small molecule free energy calculations in Q | Journal of Cheminformatics | Full Text
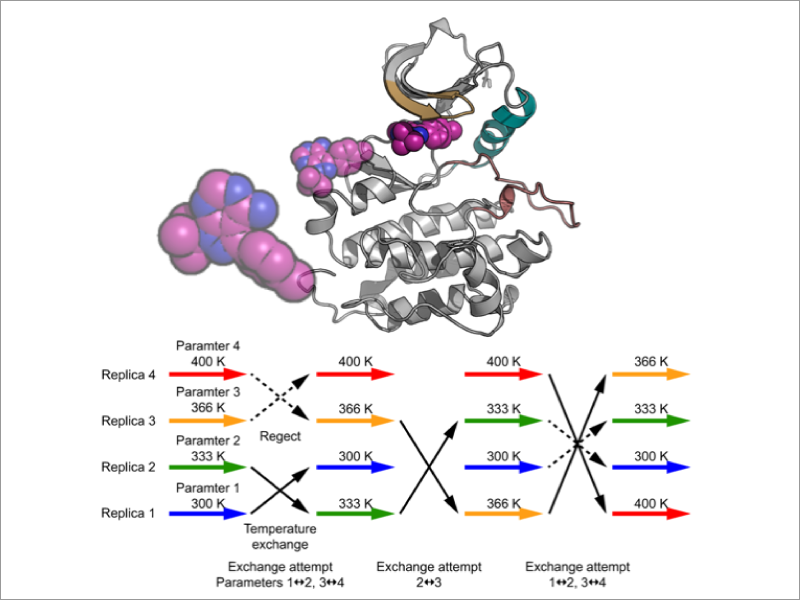
Development of free energy calculation methods for biomolecular associations – Laboratory for Biomolecular Function Simulation

PyAutoFEP: An Automated Free Energy Perturbation Workflow for GROMACS Integrating Enhanced Sampling Methods | Journal of Chemical Theory and Computation
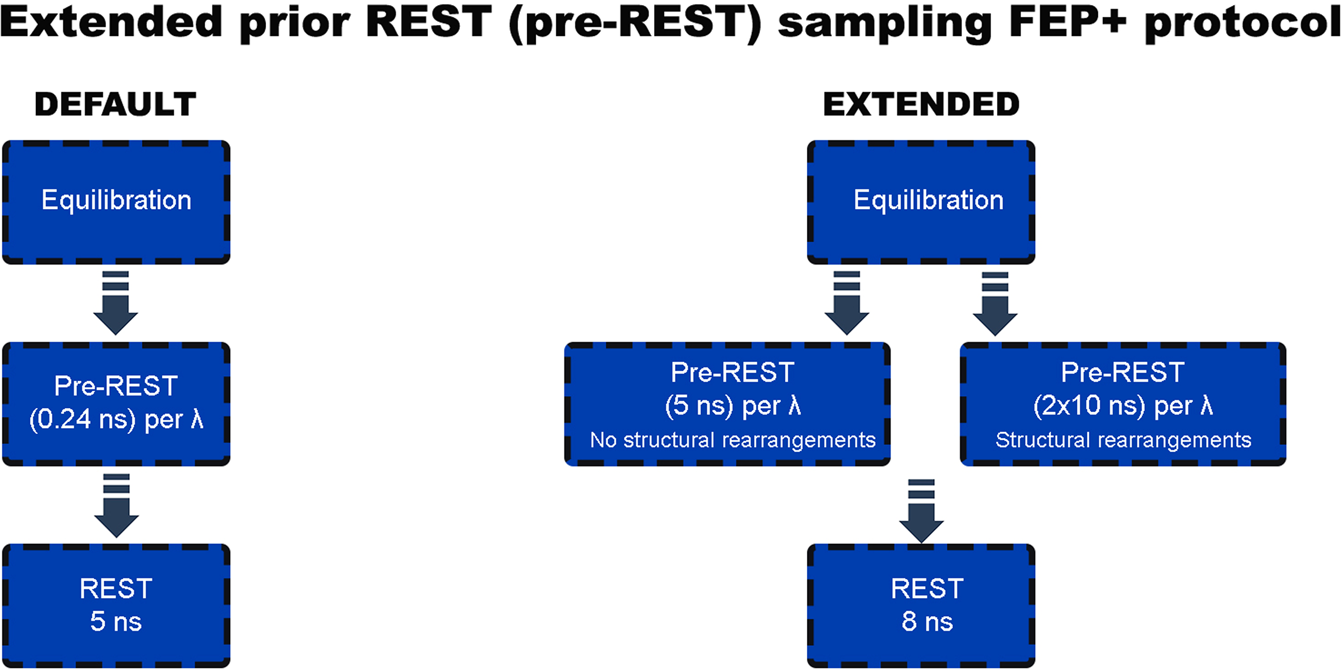
An Improved Free Energy Perturbation FEP+ Sampling Protocol for Flexible Ligand-Binding Domains | Scientific Reports

Free energy calculations General methods Free energy is the most important quantity that characterizes a dynamical process. Two types of free energy calculations: - ppt download

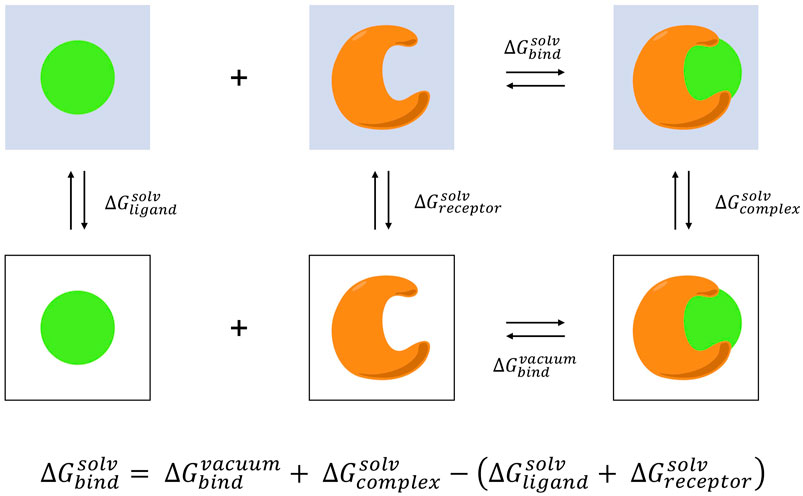
Description of the FEP calculations. A , thermodynamic cycle for FEP... | Download Scientific Diagram

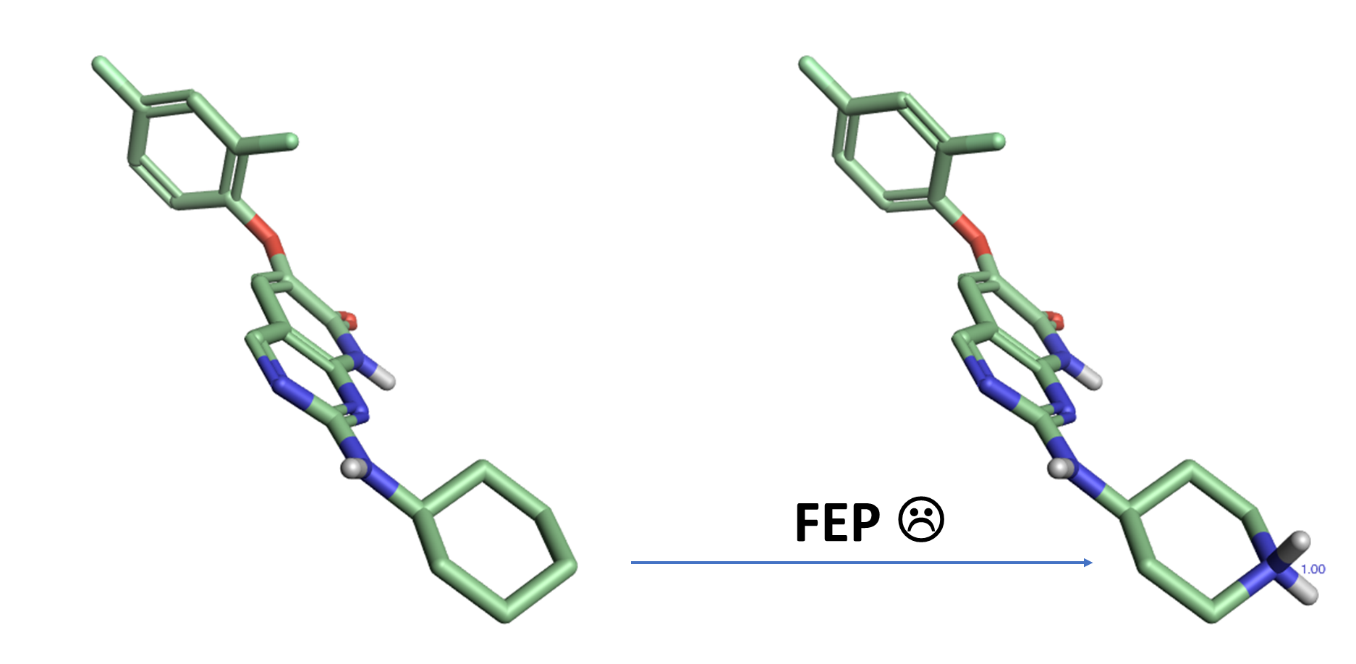
An example of FEP calculation on core-hopping using fragment-based design. | Download Scientific Diagram

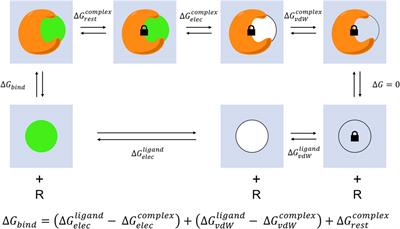
Absolute Binding Free Energy Calculation and Design of a Subnanomolar Inhibitor of Phosphodiesterase-10 | Journal of Medicinal Chemistry

IJMS | Free Full-Text | Absolute Binding Free Energy Calculations for Highly Flexible Protein MDM2 and Its Inhibitors
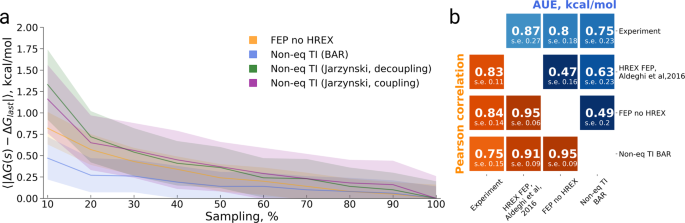
Accurate absolute free energies for ligand–protein binding based on non-equilibrium approaches | Communications Chemistry