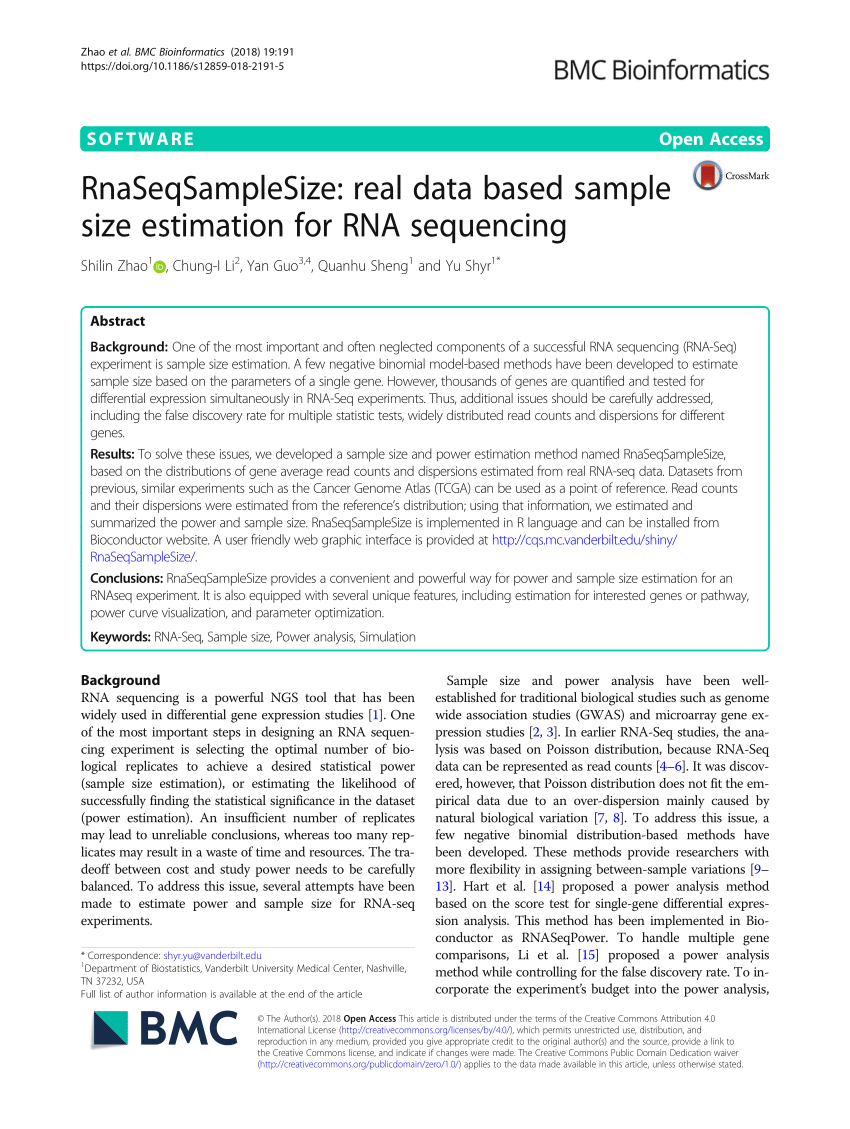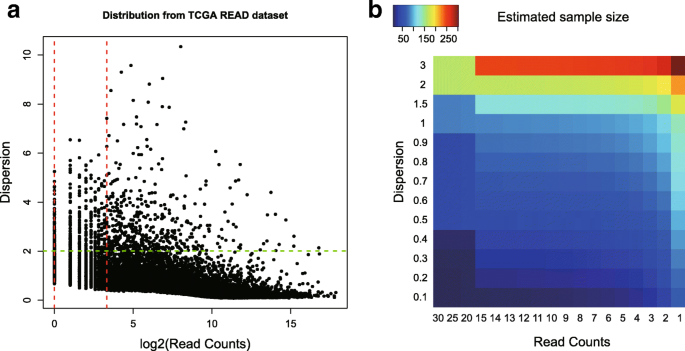
RnaSeqSampleSize: real data based sample size estimation for RNA sequencing | BMC Bioinformatics | Full Text
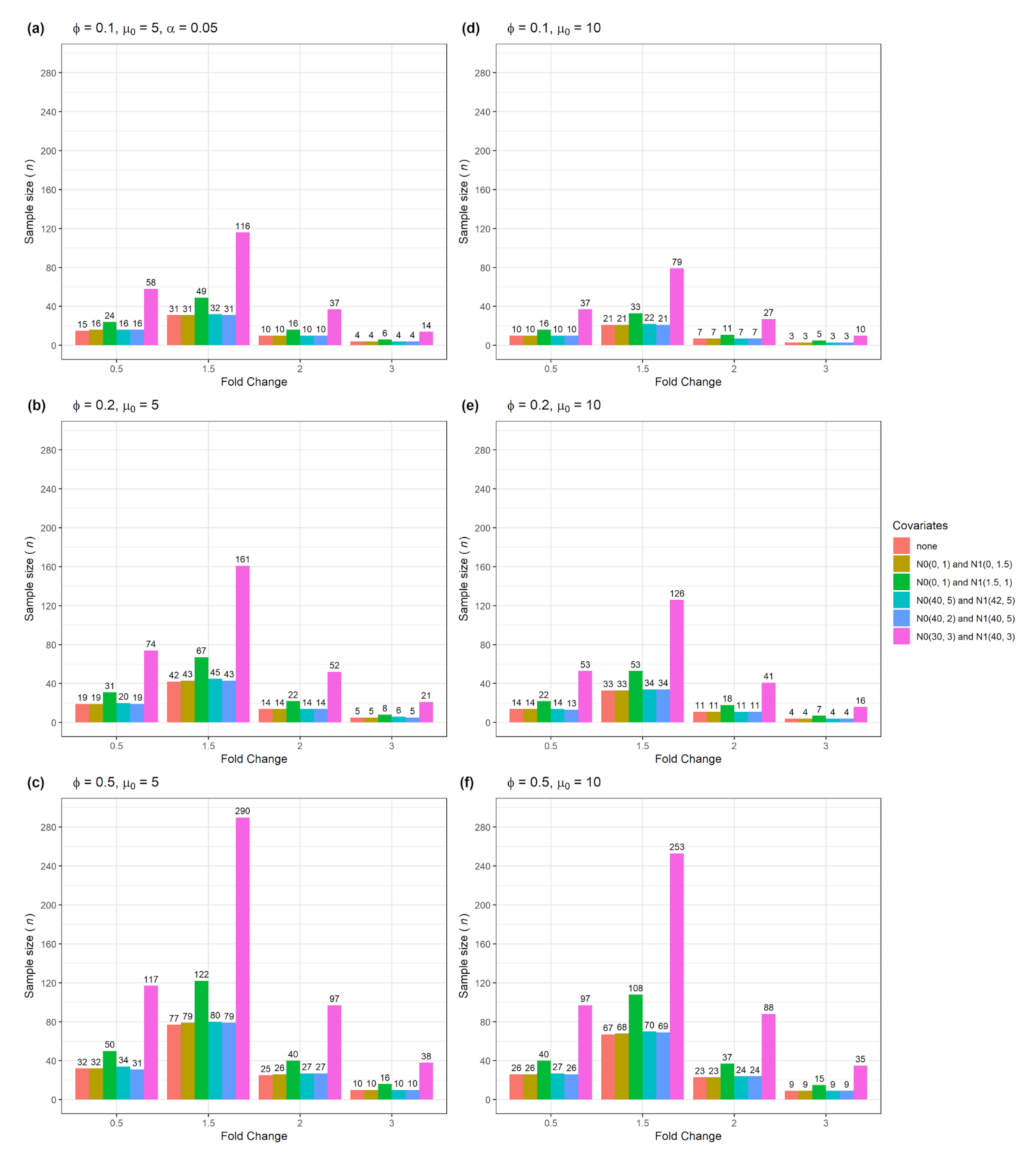
BioMedInformatics | Free Full-Text | Adjusted Sample Size Calculation for RNA-seq Data in the Presence of Confounding Covariates
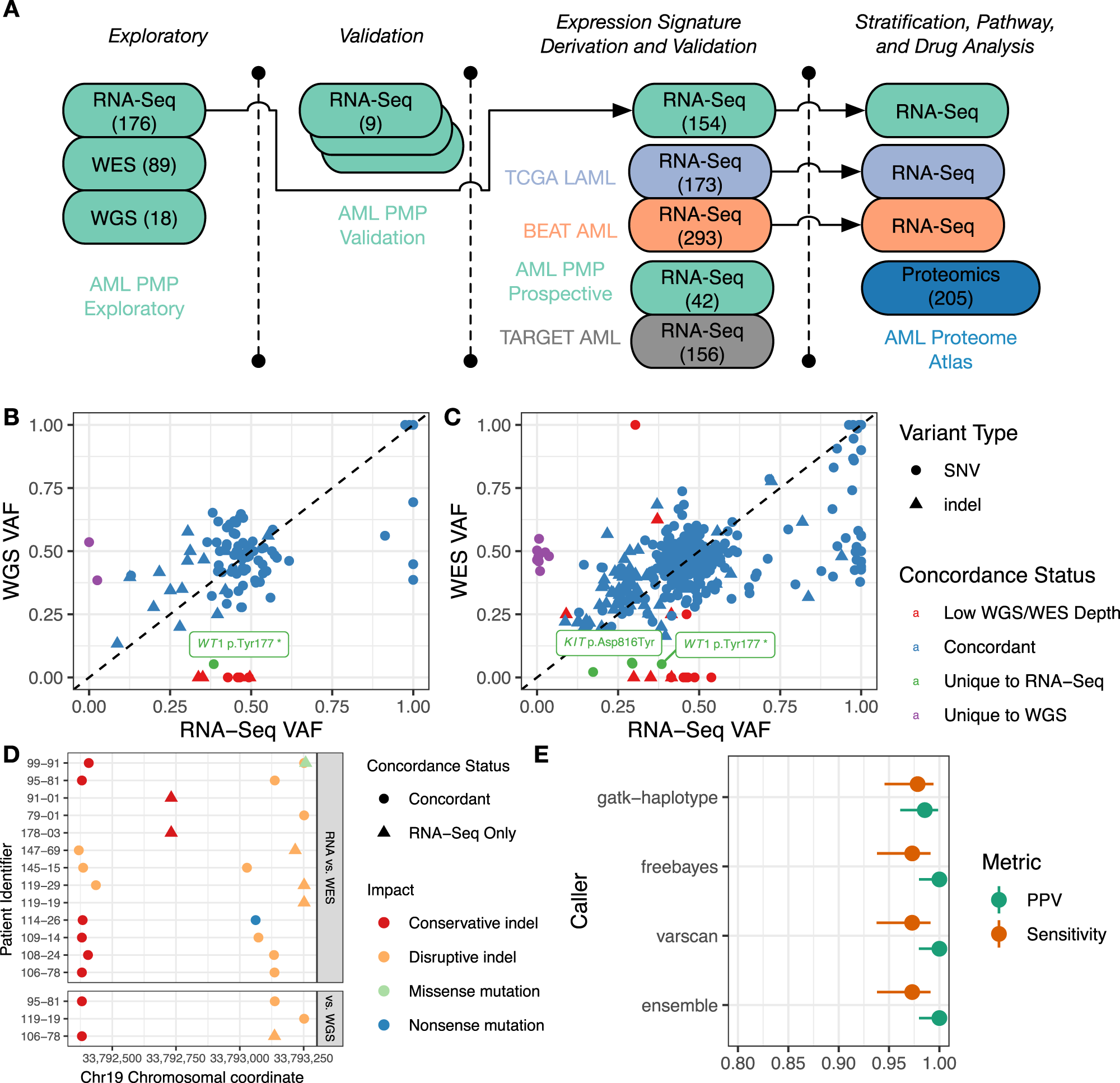
A clinical transcriptome approach to patient stratification and therapy selection in acute myeloid leukemia | Nature Communications
![PDF] Sample Size Calculation of RNA-sequencing Experiment-A Simulation-Based Approach of TCGA Data | Semantic Scholar PDF] Sample Size Calculation of RNA-sequencing Experiment-A Simulation-Based Approach of TCGA Data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6be64ffc47a1a40f43f92f648eccba9c1dae1648/4-Figure3-1.png)
PDF] Sample Size Calculation of RNA-sequencing Experiment-A Simulation-Based Approach of TCGA Data | Semantic Scholar
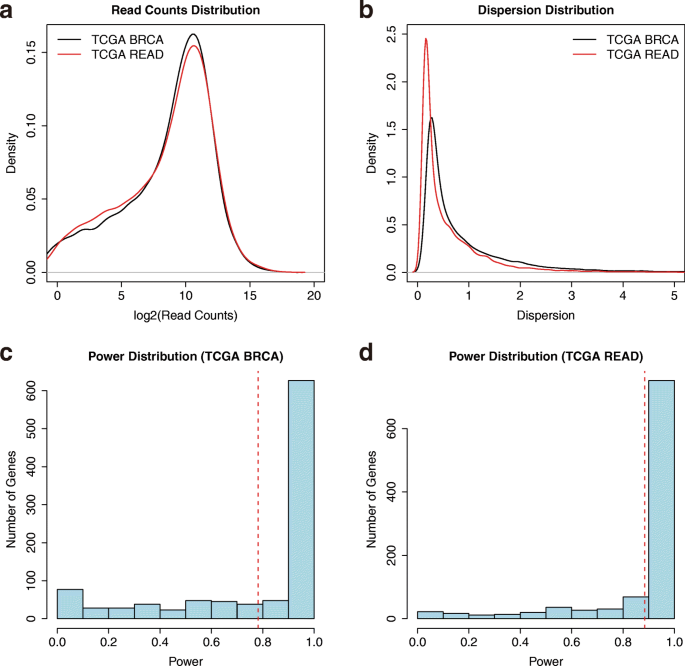
RnaSeqSampleSize: real data based sample size estimation for RNA sequencing | BMC Bioinformatics | Full Text

BioMedInformatics | Free Full-Text | Adjusted Sample Size Calculation for RNA-seq Data in the Presence of Confounding Covariates
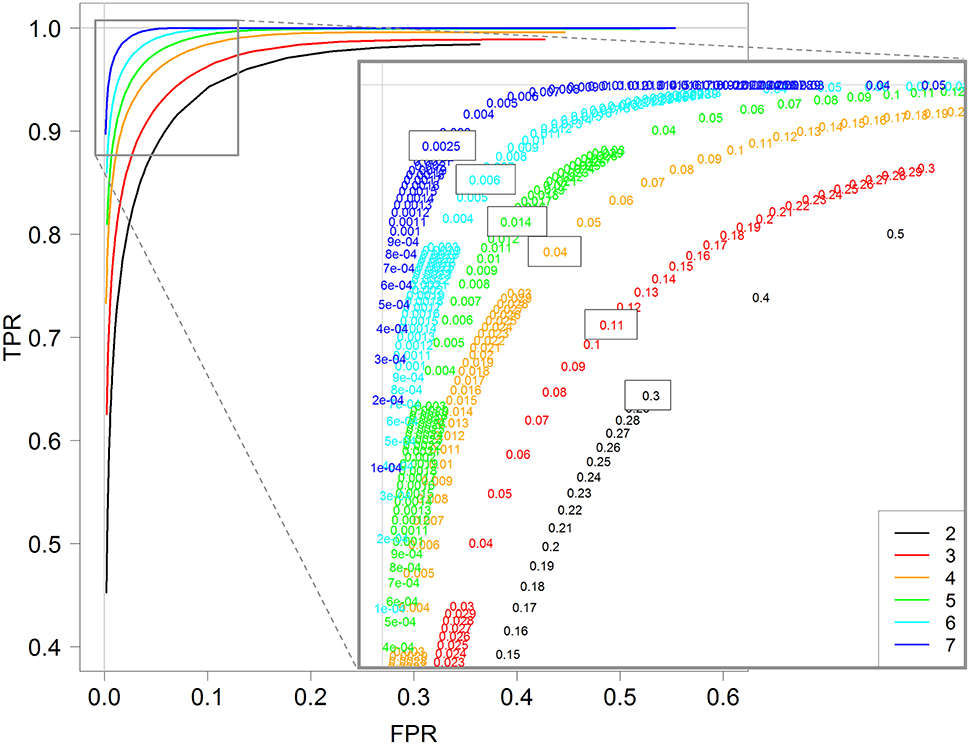
Frontiers | Optimization of an RNA-Seq Differential Gene Expression Analysis Depending on Biological Replicate Number and Library Size
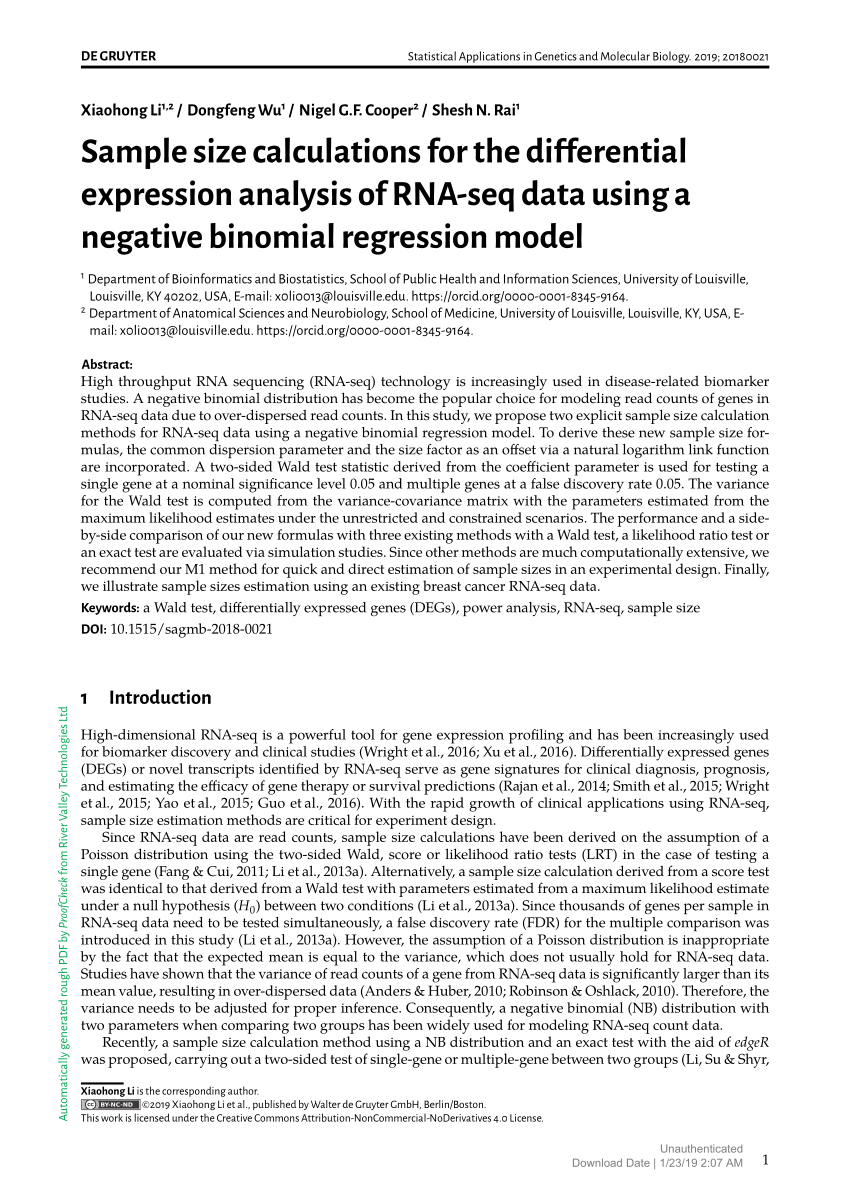

PDF) Sample size calculations for the differential expression analysis of RNA-seq data using a negative binomial regression model
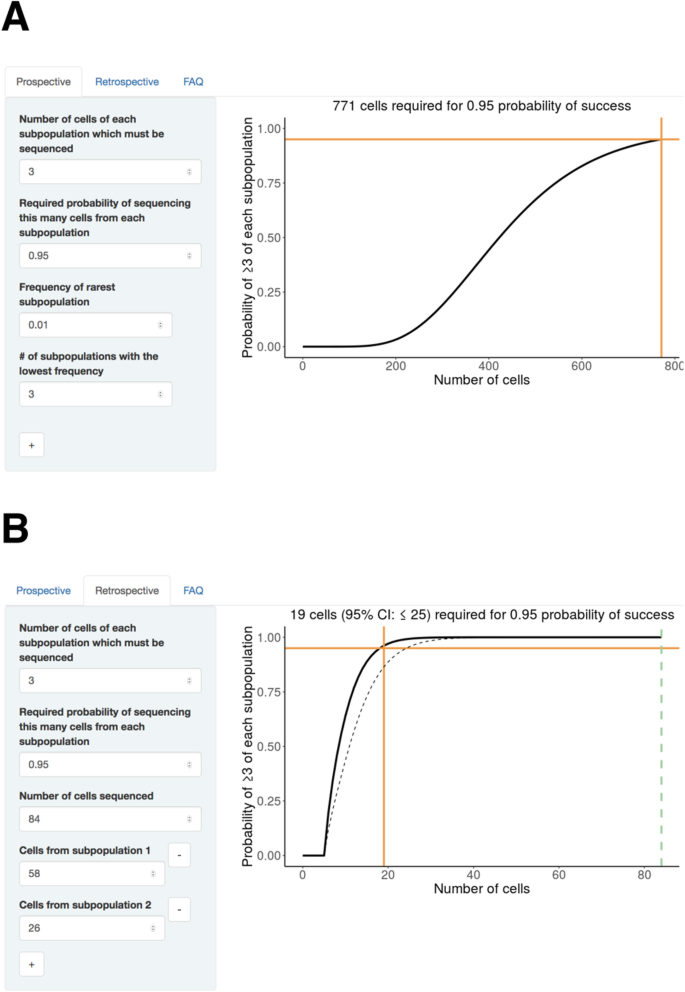
SCOPIT: sample size calculations for single-cell sequencing experiments | BMC Bioinformatics | Full Text
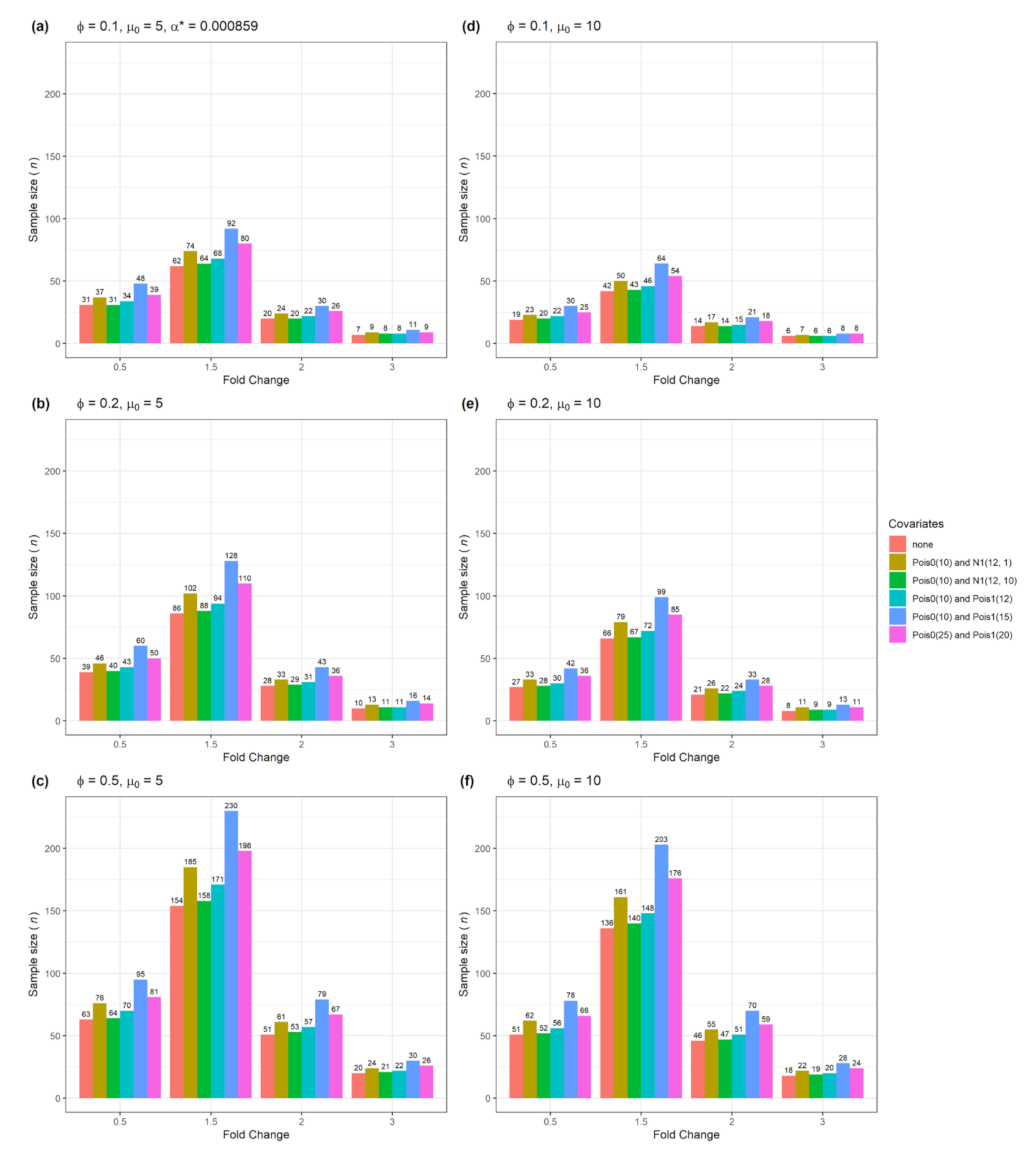
BioMedInformatics | Free Full-Text | Adjusted Sample Size Calculation for RNA-seq Data in the Presence of Confounding Covariates

Power and sample size calculations for high-throughput sequencing-based experiments | Semantic Scholar
![PDF] Sample Size Calculation of RNA-sequencing Experiment-A Simulation-Based Approach of TCGA Data | Semantic Scholar PDF] Sample Size Calculation of RNA-sequencing Experiment-A Simulation-Based Approach of TCGA Data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6be64ffc47a1a40f43f92f648eccba9c1dae1648/3-Figure1-1.png)



![PDF] Calculating Sample Size Estimates for RNA Sequencing Data | Semantic Scholar PDF] Calculating Sample Size Estimates for RNA Sequencing Data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/c1ea7e8c8542a305c2f1c771e0320bb9b578eeb4/3-Table1-1.png)


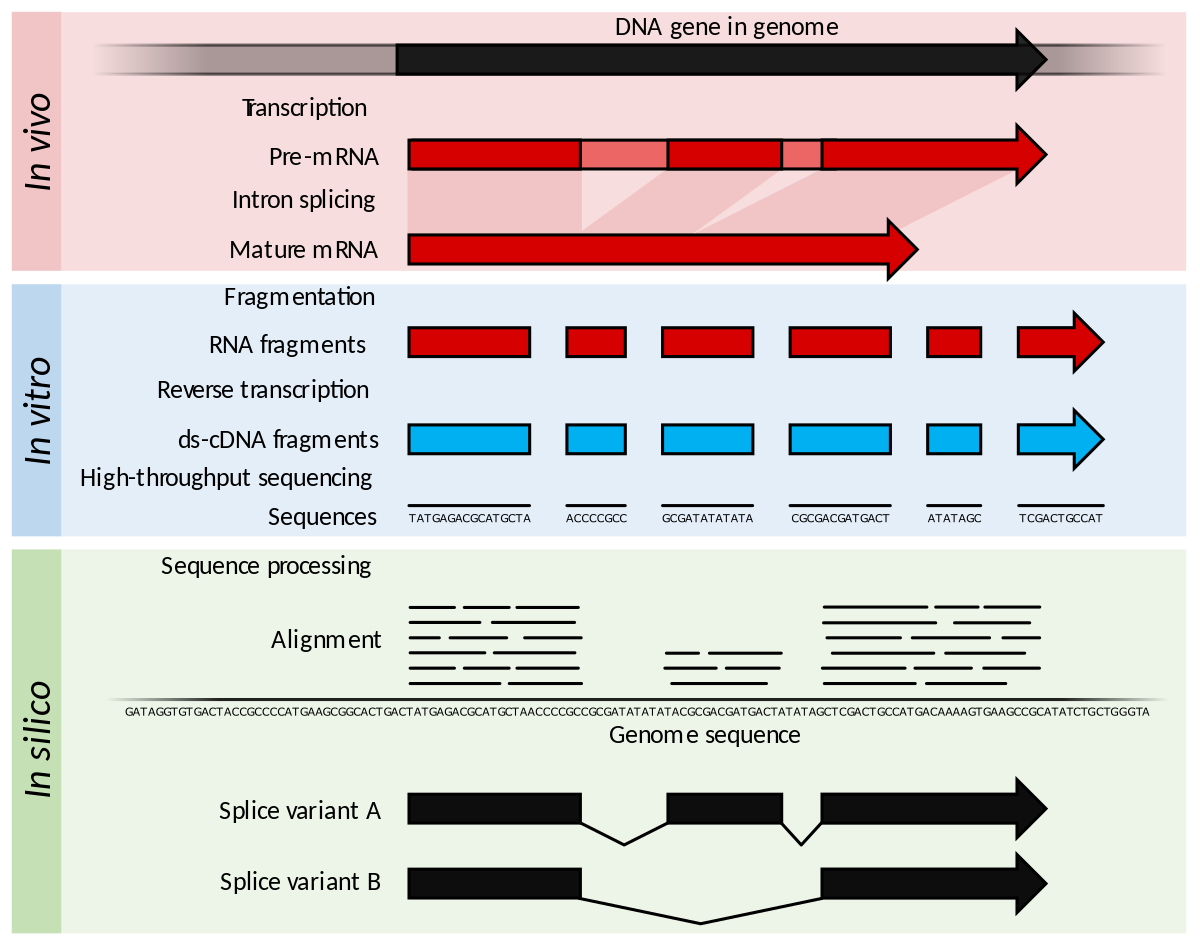
![PDF] Calculating Sample Size Estimates for RNA Sequencing Data | Semantic Scholar PDF] Calculating Sample Size Estimates for RNA Sequencing Data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/c1ea7e8c8542a305c2f1c771e0320bb9b578eeb4/6-Figure4-1.png)

